Abstract
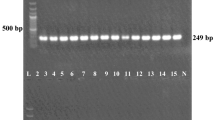
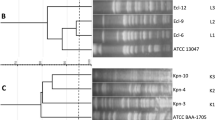
Monitoring the emergence and transmission of Pseudomonas aeruginosa strains is important for infection control worldwide. The aim of this study is to determine the genetic diversity of 47 extended-spectrum beta-lactamase-producing P. aeruginosa strains of clinical origin by enterobacterial repetitive intergenic consensus PCR (ERIC-PCR) and also to ascertain the presence of class 1 integrons among them. ERIC sequences are 127 bp imperfect palindromes that occur in multiple copies in the genomes of enteric bacteria. The ERIC-PCR profile generated showed polymorphism in the studied strains. Based on the ERIC-PCR profile, P. aeruginosa strains were grouped into 23 types. The major genotype, designated as type L1, accounted for 10.6% (5/47) of samples. Class 1 integrons were detected in 8 of the 47 clinical isolates. Further, the integron-carrying isolates were found to display higher minimum inhibitory concentration against tested antibiotics. Optimization of antimicrobial use and control of infection is recommended to prevent the increase in the population of drug-resistant organisms.



Similar content being viewed by others
References
Ahmed MU, Dunn L, Ivanova EP (2012) Evaluation of current molecular approaches for genotyping of Campylobacter jejuni strains. Food Borne Pathog. Dis 9:375–385
Chang CY, Fang YT, Tsai SM, Chang LL, Yu WL (2009) Characterization of class 1 integrons and gene cassettes in clinical isolates of Klebsiella pneumoniae from Taiwan. Diagn Microbiol Infect Dis 65:214–216
Chen J, Su Z, Liu Y, Wang S, Dai X, Li Y, Peng S, Shao Q, Zhang H, Wen P, Yu J, Huang X, Xu H (2009) Identification and characterization of class 1 integrons among Pseudomonas aeruginosa isolates from patients in Zhenjiang, China. Int J Infect Dis 13:717–721
Clinical and Laboratory Standards Institute (2008) Performance standards for antimicrobial susceptibility testing: eighteenth informational supplement. Wayne, PA
Fonseca EL, Vieira VV, Cipriano R, Vicente AC (2005) Class 1 integrons in Pseudomonas aeruginosa isolates from clinical settings in Amazon region, Brazil. FEMS Immunol Med Microbiol 44:303–309
Gu B, Tong M, Zhao W, Liu G, Ning M, Pan S (2007) Prevalence and characterization of class I integrons among Pseudomonas aeruginosa and Acinetobacter baumannii isolates from patients in Nanjing China. J Clin Microbiol 45:241–243
Guerra B, Soto S, Cal S, Mendoza MC (2000) Antimicrobial resistance and spread of class 1 integrons among Salmonella Serotypes. Antimicrob Agents Chemother 44:2166–2169
Gutirrez O, Juan C, Cercenado E, Navarro F, Bouza E, Coll P, Perez JL, Oliver A (2007) Molecular epidemiology and mechanisms of carbapenem resistance in Pseudomonas aeruginosa isolates from Spanish hospitals. Antimicrob Agents Chemother 51:4329–4335
Juan C, Beceiro A, Gutierrez O, Alberti S, Garau M, Perez JL, Bou G, Oliver A (2008) Characterization of the new metallo-beta-lactamase VIM-13 and its integron-borne gene from a Pseudomonas aeruginosa clinical isolate in Spain. Antimicrob Agents Chemother 52:3589–3596
Kidd TJ, Gibson JS, Moss S, Greer RM, Cobbold RN, Wright JD, Grimwood K, Bell SC (2011) Clonal complex Pseudomonas aeruginosa in horses. Vet Microbiol 149:508–512
Lang XY, Zhang YL, Jiang HT, Liu J (2013) Development of an enterobacterial repetitive intergenic consensus polymerase chain reaction (ERIC-PCR) to detect and genotype enterotoxigenic Escherichia coli of calf origin. Afr J Microbiol Res 7:4001–4005
Li X, Shi L, Yang W, Li L, Yamasaki S (2006) New array of aacA4- catB3 dfrA1 gene cassettes and a noncoding cassette from a class-1-integron-positive clinical strain of Pseudomonas aeruginosa. Antimicrob Agents Chemother 50:2278–2279
Li W, Raoult D, Fournier PE (2009) Bacterial strain typing in genomic era. FEMS Microbiol Rev 33:892–916
Liu Y, Davin-Regli A, Bosi C, Charrel RN, Bollet C (1996) Epidemiological investigation of Pseudomonas aeruginosa nosocomial bacteraemia isolates by PCR-based DNA fingerprinting analysis. J Med Microbiol 45:359–365
Macedo NR, Oliceira SR, Lage AP, Sanros JL, Araújo MR, Guedes RMC (2011) ERIC-PCR genotyping of Haemophilus parasuis isolates from Brazilian pigs. Vet J 188:362–364
Pitout JDD, Chow BL, Gregson DB, Laupland KB, Elsayed S, Church DL (2007) Molecular epidemiology of metallo-beta-lactamase-producing Pseudomonas aeruginosa in the Calgary health region: emergence of VIM-2-producing isolates. J Clin Microbiol 45:294–298
Rajabnia R, Asgharpour F, Ferdosi Shahandashti E, Khalilian M, Norkhomami S, Shafii M, Moulana Z (2013) Class 1 Integron in Pseudomonas aeruginosa isolates from different places and devices of ICU in Babol, Iran. Jundishapur J Microbiol 6:138–143
Rao AN, Barlow M, Clark LA, Boring JR, Tenover FC, McGowan JE (2006) Class 1 integrons in resistant Escherichia coli and Klebsiella spp., US hospitals. Emerg Infect Dis 12:1011–1014
Ruiz-Martinez L, Lopez-Jimenez L, Fuste E, Vinuesa T, Martinez JP, Vinas M (2011) Class 1 integrons in environmental and clinical isolates of Pseudomonas aeruginosa. Int J Antimicrob Agents 38:398–402
Shi L, Zheng M, Xiao Z, Asakura M, Su J, Li L, Yamasaki S (2006) Unnoticed spread of class 1 integrons in gram-positive strains isolated in Guangzhou, China. Microbiol Immunol 50:463–467
Speijer H, Savelkoul PH, Bonten MJ, Stobberingh EE, Tjhie JH (1999) Application of different genotyping methods for Pseudomonas aeruginosa in a setting of endemicity in an intensive care unit. J Clin Microbiol 37:3654–3661
Stehling EG, Leite DS, Sileira WD (2010) Molecular typing and biological characteristics of Pseudomonas aeruginosa isolated from cystic fibrosis patients in Brazil. Braz J Infect Dis 14:462–467
Tazumi A, Maeda Y, Buckley T, Millar BC, Goldsmith CE, Dooley JSG, Eldorn JS, Matsuda M, Moore JE (2009) Molecular epidemiology of clinical isolates of Pseudomonas aeruginosa isolated from horses in Ireland. Ir Vet J 62:456–459
Versalovic J, Koeuth T, Lupskil JR (1991) Distribution of repetitive DNA sequences in eubacteria and application to fingerprinting of bacterial genomes. Nucleic Acids Res 19:6823–6831
Wang CY, Jerng JS, Chen KY, Lee LN, Yu CJ, Hsueh PR, Yang PC (2006) Pandrug-resistant Pseudomonas aeruginosa among hospitalized patients: clinical features, risk-factors and outcomes. Clin Microbiol Infect 12:63–68
Xu Z, Shi L, Zhang C, Zhang L, Li X, Cao Y, Li L, Yamasaki S (2007) Nosocomial infection caused by class 1 integron-carrying Staphylococcus aureus in a hospital in South China. Clin Microbiol Infect 13:980–984
Xu Z, Li L, Alam MJ, Yamasaki S, Shi L (2008) First confirmation of integron-bearing methicillin-resistant Staphylococcus aureus. Curr Microbiol 57:264–268
Acknowledgements
Shaikh S was supported by INSPIRE grant from the Department of Science and Technology, New Delhi, India (Grant Number: IF130056), which is sincerely acknowledged. Shazi Shakil thanks all of the staff of KACST Technology Innovation Center for Personalized Medicine at King Abdulaziz University, Jeddah, Saudi Arabia, for continued support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Shaikh, S., Rizvi, S.M.D., Shakil, S. et al. Non-clonal Dissemination of Extended-Spectrum Beta-Lactamase-Producing Pseudomonas aeruginosa Strains of Clinical Origin. Iran J Sci Technol Trans Sci 41, 1011–1015 (2017). https://doi.org/10.1007/s40995-017-0340-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40995-017-0340-8