Abstract
Introduction
Different normalization methods are available for urinary data. However, it is unclear which method performs best in minimizing error variance on a certain data-set as no generally applicable empirical criteria have been established so far.
Objectives
The main aim of this study was to develop an applicable and formally correct algorithm to decide on the normalization method without using phenotypic information.
Methods
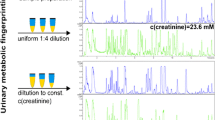
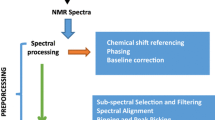
We proved mathematically for two classical measurement error models that the optimal normalization method generates the highest correlation between the normalized urinary metabolite concentrations and its blood concentrations or, respectively, its raw urinary concentrations. We then applied the two criteria to the urinary 1H-NMR measured metabolomic data from the Study of Health in Pomerania (SHIP-0; n = 4068) under different normalization approaches and compared the results with in silico experiments to explore the effects of inflated error variance in the dilution estimation.
Results
In SHIP-0, we demonstrated consistently that probabilistic quotient normalization based on aligned spectra outperforms all other tested normalization methods. Creatinine normalization performed worst, while for unaligned data integral normalization seemed to most reasonable. The simulated and the actual data were in line with the theoretical modeling, underlining the general validity of the proposed criteria.
Conclusions
The problem of choosing the best normalization procedure for a certain data-set can be solved empirically. Thus, we recommend applying different normalization procedures to the data and comparing their performances via the statistical methodology explicated in this work. On the basis of classical measurement error models, the proposed algorithm will find the optimal normalization method.

Similar content being viewed by others
References
Alonso, A., Rodríguez, M. A., Vinaixa, M., Tortosa, R., Correig, X., Julià, A., & Marsal, S. (2014). Focus: A robust workflow for one-dimensional NMR spectral analysis. Analytical Chemistry, 86(2), 1160–1169. doi:10.1021/ac403110u.
Bouatra, S., Aziat, F., Mandal, R., Guo, A. C., Wilson, M. R., Knox, C., et al. (2013). The human urine metabolome. PLoS ONE, 8(9), e73076. doi:10.1371/journal.pone.0073076.
Buonaccorsi, J. P. (2010). Measurement error: Models, methods, and applications. Boca Raton: CRC Press.
Chadha, V., Alon, U. S., & Garg, U. (2001). Measurement of urinary concentration: A critical appraisal of methodologies. Pediatric Nephrology, 16(4), 374–382. doi:10.1007/s004670000551.
Cooke, D. W., & Plotnick, L. (2008). Type 1 diabetes mellitus in pediatrics. Pediatrics in Review, 29(11), 374–384 quiz 385. doi:10.1542/pir.29-11-374.
Dieterle, F., Ross, A., Schlotterbeck, G., & Senn, H. (2006). Probabilistic quotient normalization as robust method to account for dilution of complex biological mixtures. Application in 1H NMR metabonomics. Analytical Chemistry, 78(13), 4281–4290. doi:10.1021/ac051632c.
Duarte, I. F., Diaz, S. O., & Gil, A. M. (2014). NMR metabolomics of human blood and urine in disease research. Journal of Pharmaceutical and Biomedical Analysis, 93, 17–26. doi:10.1016/j.jpba.2013.09.025.
Emwas, A.-H., Luchinat, C., Turano, P., Tenori, L., Roy, R., Salek, R. M., et al. (2015). Standardizing the experimental conditions for using urine in NMR-based metabolomic studies with a particular focus on diagnostic studies: a review. Metabolomics, 11(4), 872–894. doi:10.1007/s11306-014-0746-7.
González-Domínguez, R., Castilla-Quintero, R., García-Barrera, T., & Gómez-Ariza, J. L. (2014). Development of a metabolomic approach based on urine samples and direct infusion mass spectrometry. Analytical Biochemistry, 465, 20–27. doi:10.1016/j.ab.2014.07.016.
Hertel, J., Friedrich, N., Wittfeld, K., Pietzner, M., Budde, K., Van der Auwera, S., et al. (2016). Measuring Biological Age via Metabonomics: The Metabolic Age Score. Journal of Proteome Research, 15(2), 400–410. doi:10.1021/acs.jproteome.5b00561.
Kohl, S. M., Klein, M. S., Hochrein, J., Oefner, P. J., Spang, R., & Gronwald, W. (2012). State-of-the art data normalization methods improve NMR-based metabolomic analysis. Metabolomics, 8(S1), 146–160. doi:10.1007/s11306-011-0350-z.
Rose, B. D., & Rennke, H. G. (1994). Renal pathophysiology: The essentials. Baltimore: Williams & Wilkins.
Sauvé, J.-F., Lévesque, M., Huard, M., Drolet, D., Lavoué, J., Tardif, R., & Truchon, G. (2015). Creatinine and specific gravity normalization in biological monitoring of occupational exposures. Journal of Occupational and Environmental Hygiene, 12(2), 123–129. doi:10.1080/15459624.2014.955179.
Schnackenberg, L. K., Sun, J., Espandiari, P., Holland, R. D., Hanig, J., & Beger, R. D. (2007). Metabonomics evaluations of age-related changes in the urinary compositions of male Sprague Dawley rats and effects of data normalization methods on statistical and quantitative analysis. BMC Bioinformatics, 8(Suppl 7), S3. doi:10.1186/1471-2105-8-S7-S3.
Sugimoto, M., Kawakami, M., Robert, M., Soga, T., & Tomita, M. (2012). Bioinformatics Tools for Mass Spectroscopy-Based Metabolomic Data Processing and Analysis. Current Bioinformatics, 7(1), 96–108. doi:10.2174/157489312799304431.
Völzke, H., Alte, D., Schmidt, C. O., Radke, D., Lorbeer, R., Friedrich, N., et al. (2011). Cohort profile: the study of health in Pomerania. International Journal of Epidemiology, 40(2), 294–307. doi:10.1093/ije/dyp394.
Vu, T. N., & Laukens, K. (2013). Getting your peaks in line: a review of alignment methods for NMR spectral data. Metabolites, 3(2), 259–276. doi:10.3390/metabo3020259.
Warrack, B. M., Hnatyshyn, S., Ott, K.-H., Reily, M. D., Sanders, M., Zhang, H., & Drexler, D. M. (2009). Normalization strategies for metabonomic analysis of urine samples. Journal of Chromatography B, 877(5–6), 547–552. doi:10.1016/j.jchromb.2009.01.007.
Acknowledgements
The contribution to data collection performed by study nurses, study physicians, interviewers, and laboratory workers is gratefully acknowledged. We are also appreciative of the important support of IT and computer scientists, health information managers and administration staff. We also thank all study participants whose personal dedication and commitment made this project possible.
Author contributions
JH wrote the manuscript, derived the statistical criteria and analyzed the data. MP, NF and KB provided NMR-data and data-preparation and reviewed the manuscript. KW, SVA and HJG took part in every step of the design of the manuscript. AT reviewed the mathematical derivations and the manuscript. MN and TK contributed to the design and data acquisition of the SHIP-study and MN organized the laboratory infrastructure for NMR-measurements and all laboratory work.
Funding
The SHIP study (author MN) is part of the Community Medicine Research net of the University of Greifswald, Germany, which is funded by the Federal Ministry of Education and Research (Grants no. 01ZZ9603, 01ZZ0103, and 01ZZ0403) and add-ons to this grants of the Ministry of Cultural Affairs and the Social Ministry of the Federal State of Mecklenburg-West Pomerania. This work was also part of the project GANI_MED (Greifswald Approach to Individualized Medicine, authors JH, MP, SVA, MN and HJG) which is funded by the Federal Ministry of Education and Research (Grant no. 03IS2061A).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
We have no conflict of interest to declare.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Matthias Nauck and Hans Jörgen Grabe have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hertel, J., Van der Auwera, S., Friedrich, N. et al. Two statistical criteria to choose the method for dilution correction in metabolomic urine measurements. Metabolomics 13, 42 (2017). https://doi.org/10.1007/s11306-017-1177-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-017-1177-z