Abstract
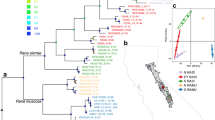
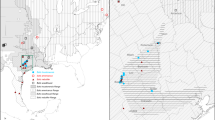
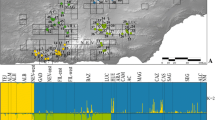
The mountain yellow-legged species complex (Rana sierrae and Rana muscosa) has declined precipitously in distribution and abundance during the last century. The two primary threats are chytrid epidemic-associated population collapses and predation from the introduction of non-native trout. Widespread declines have occurred throughout the ranges of these species, including populations of R. sierrae in Yosemite National Park. A clear picture of genetic structure of remaining Yosemite R. sierrae populations is critical to short-term management and conservation. We conducted a population genetics study that included samples from 23 geographic sites distributed throughout the range of R. sierrae in Yosemite NP. We used minimally-invasive swab samples to collect genetic data from mitochondrial and nuclear DNA via sequencing (43 transcriptome-derived markers) and analyzed the distribution of genetic variation in a geographic context. Our mtDNA analysis partially confirmed previous results suggesting that two haplotype groups occur in Yosemite: one haplotype group contained high bootstrap support for monophyly while the other did not. However, increased geographic sampling demonstrated that the two haplotypes are not completely geographically partitioned into the two main drainages (Merced drainage and Tuolumne drainage) as previously postulated. Our nuclear DNA analysis revealed a general pattern of genetic isolation by distance, where genetic differentiation was correlated with geographic distance between sites. In addition, our analyses suggested that three clusters of genetically cohesive sites occur in the study area. Understanding population genetic patterns of variability will inform management strategies such as translocations, reintroductions, and monitoring for this endangered frog. Lastly, our next generation sequencing enabled approach allowed us to obtain multi-locus data from minimally-invasive swab samples. Thus researchers can now leverage extensive archives of swab samples (initially collected for pathogen testing) to study host genetics in previously surveyed amphibian populations.







Similar content being viewed by others
References
Adamack AT, Gruber B (2014) PopGenReport: simplifying basic population genetic analyses in R. Methods Ecol Evol 5:384–387. doi:10.1111/2041-210X.12158
Blouin MS, Phillipsen IC, Monsen KJ (2010) Population structure and conservation genetics of the Oregon spotted frog, Rana pretiosa. Conserv Genet (2010) 11: 2179–2194. doi:10.1007/s10592-010-0104-x
Briggs CJ, Knapp RA, Vredenburg VT (2010) Enzootic and epizootic dynamics of the chytrid fungal pathogen of amphibians. Proc Natl Acad Sci USA 107:9695–9700
Chevreux B, Pfisterer T, Drescher B et al (2004) Using the miraEST assembler for reliable and automated mRNA transcript assembly and SNP detection in sequenced ESTs. Genome Res 14:1147–1159
Cushman S, Wasserman T, Landguth E, Shirk A (2013) Re-evaluating causal modeling with mantel tests in landscape genetics. Diversity 5:51–72
Dellicour S, Mardulyn P (2013) spads 1.0: A toolbox to perform spatial analyses on DNA sequence data sets. Mol Ecol Resour 14:647–651
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797. doi:10.1093/nar/gkh340
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Grinnell J, Storer T (1924) Animal life in the yosemite. University of California Press, Berkeley
Gruber B, Adamack AT (2015) Landgenreport: a new R function to simplify landscape genetic analysis using resistance surface layers. Mol. Ecol Res 15:1172–1178
Guillot G, Leblois R, Coulon A, Frantz AC (2009) Statistical methods in spatial genetics. Mol Ecol 18:4734–4756
Hyatt AD, Boyle DG, Olsen V et al (2007) Diagnostic assays and sampling protocols for the detection of Batrachochytrium dendrobatidis. Dis Aquat Organ 73:175–192
Jombart T (2008) adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24:1403–1405. doi:10.1093/bioinformatics/btn129
Jombart T, Devillard S, Dufour A-B, Pontier D (2008) Revealing cryptic spatial patterns in genetic variability by a new multivariate method. Heredity 101:92–103
Kalinowski ST (2009) How well do evolutionary trees describe genetic relationships among population. Heredity 102:506–513
Kearse M, Moir R, Wilson A et al (2012) Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649
Knapp RA (2005) Effects of nonnative fish and habitat characteristics on lentic herpetofauna in Yosemite National Park, USA. Biol Conserv 121:265–279. doi:10.1016/j.biocon.2004.05.003
Knapp RA, Matthews KR (2000) Non-native fish introductions and the decline of the mountain yellow-legged frog from within protected areas. Conserv Biol 14:428–438
Knapp RA, Briggs CJ, Smith TC, Maurer JR (2011) Nowhere to hide: impact of a temperature-sensitive amphibian pathogen along an elevation gradient in the temperate zone. Ecosphere 2:1–26
Kopelman NM, Mayzel J, Jakobsson M, et al. (2015) Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Mol Ecol Resour 15: 1179–1191. doi:10.1111/1755-0998.12387
Legendre P, Fortin MJ (2010) Comparison of the Mantel test and alternative approaches for detecting complex multivariate relationships in the spatial analysis of genetic data. Mol. Ecol Res 10:831–844. doi:10.1111/j.1755-0998.2010.02866.x
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760. doi:10.1093/bioinformatics/btp324
Lind AJ, Spinks PQ, Fellers GM, Shaffer HB (2011) Rangewide phylogeography and landscape genetics of the Western US endemic frogRana boylii(Ranidae): implications for the conservation of frogs and rivers. Conserv Genet 12:269–284
Magoc T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27:2957–2963. doi:10.1093/bioinformatics/btr507
McCartney - Melstad E, Shaffer HB (2015) Amphibian molecular ecology and how it has informed conservation. Mol Ecol 24:5084–5109
Monsen KJ, Blouin MS (2003) Genetic structure in a montane ranid frog: restricted gene flow and nuclear-mitochondrial discordance. Mol Ecol 12:3275–3286
Monsen KJ, Blouin MS (2004) Extreme isolation by distance in a montane frog Rana cascadae. Conserv Genet 5:827–835. doi:10.1007/s10592-004-1981-z
Novembre J, Johnson T, Bryc K, et al. (2008) Genes mirror geography within Europe. Nature 456:98–101
Novembre J, Stephens M (2008) Interpreting principal component analyses of spatial population genetic variation. Nature Genet 40:646–649
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rosenblum EB, Poorten TJ, Settles M, Murdoch GK (2012) Only skin deep: shared genetic response to the deadly chytrid fungus in susceptible frog species. Mol Ecol 21:3110–3120. doi:10.1111/j.1365-294X.2012.05481.x
Rovito SM (2010) Lineage divergence and speciation in the Web-toed Salamanders (Plethodontidae: Hydromantes) of the Sierra Nevada, California. Mol Ecol 19:4554–4571. doi:10.1111/j.1365-294X.2010.04825.x
Schliep KP (2011) Phangorn: phylogenetic analysis in R. Bioinformatics 27:592–593. doi:10.1093/bioinformatics/btq706
Schoville SD, Roderick GK, Kavanaugh DH (2012) Testing the “Pleistocene species pump”in alpine habitats: lineage diversification of flightless ground beetles (Coleoptera: Carabidae: Nebria) in relation to altitudinal zonation. Biol J Linn Soc 107:95–111.
Semlitsch RD (2002) Critical Elements for Biologically Based Recovery Plans of Aquatic-Breeding Amphibians. Conserv Biol 16:619–629. doi:10.1046/j.1523-1739.2002.00512.x
Shaffer HB, Fellers GM, Magee A, Voss SR (2000) The genetics of amphibian declines: population substructure and molecular differentiation in the Yosemite Toad, Bufo canorus (Anura, Bufonidae) based on single-strand conformation polymorphism analysis (SSCP) and mitochondrial DNA sequence data. Mol Ecol 9:245–257. doi:10.1046/j.1365-294x.2000.00835.x
Shaffer HB, Fellers GM, Voss SR, Oliver JC, Pauly GB (2004) Species boundaries, phylogeography and conservation genetics of the red - legged frog (Rana aurora / draytonii) complex. Mol Ecol 13:2667–2677
Smedley D, Haider S, Ballester B et al (2009) BioMart–biological queries made easy. BMC Genom 10:22. doi:10.1186/1471-2164-10-22
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–2690. doi:10.1093/bioinformatics/btl446
Stuart SN, Chanson JS, Cox NA, Young BE, Rodrigues AS, Fischman DL, Waller RW (2004) Status and trends of amphibian declines and extinctions worldwide. Science 306:1783–1786
Toews DPL, Brelsford A (2012) The biogeography of mitochondrial and nuclear discordance in animals. Mol Ecol 21:3907–3930. doi:10.1111/j.1365-294X.2012.05664.x
Van der Auwera GA, Carneiro MO, Hartl C, et al. (2013) From fastQ data to high-confidence variant calls: the genome analysis toolkit best practices pipeline. Curr Protoc Bioinforma doi:10.1002/0471250953.bi1110s43
Vredenburg VT, Bingham R, Knapp R et al (2007) Concordant molecular and phenotypic data delineate new taxonomy and conservation priorities for the endangered mountain yellow-legged frog. J Zool 271:361–374. doi:10.1111/j.1469-7998.2006.00258.x
Vredenburg VT, Knapp RA, Tunstall TS, Briggs CJ (2010) Dynamics of an emerging disease drive large-scale amphibian population extinctions. Proc Natl Acad Sci USA 107:9689–9694. doi:10.1073/pnas.0914111107
Wang IJ (2013) Examining the full effects of landscape heterogeneity on spatial genetic variation: a multiple matrix regression approach for quantifying geographic and ecological isolation. Evol Int J org Evol 67:3403–3411
Zieliński P, Stuglik MT, Dudek K, et al. (2014) Development, validation and high-throughput analysis of sequence markers in nonmodel species. Mol Ecol Resour 14:352–360. doi:10.1111/1755-0998.12171
Acknowledgements
We thank Mary Toothman for assistance in retrieving archived samples, Jeanine Refsnider for assistance with experimental design, Rob Grasso for helpful comments on the manuscript, Patrick Kleeman and Brian Halstead for assistance with sample collection from the Summit Meadow frog population, and many field assistants, especially Tom Smith, Neil Kauffman, and Maxwell Joseph, for sample collection at the remaining sites. All sample collection (skin swabs) was led by Roland Knapp and authorized by research permits provided by Yosemite National Park and the Institutional Animal Care and Use Committee at University of California, Santa Barbara (UCSB). Funding was provided by the Yosemite Conservancy and a Valentine Eastern Sierra Reserve Graduate Student Research Grant from the University of California Natural Reserve System.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Poorten, T.J., Knapp, R.A. & Rosenblum, E.B. Population genetic structure of the endangered Sierra Nevada yellow-legged frog (Rana sierrae) in Yosemite National Park based on multi-locus nuclear data from swab samples. Conserv Genet 18, 731–744 (2017). https://doi.org/10.1007/s10592-016-0923-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-016-0923-5