Abstract
Purpose
Early identification of inborn errors of immunity (IEIs) is crucial due to the significant risk of morbidity and mortality. This study aimed to describe the genetic causes, clinical features, and survival rate of IEIs in Omani patients.
Methods
A prospective study of all Omani patients evaluated for immunodeficiency was conducted over a 17-year period. Clinical features and diagnostic immunological findings were recorded. Targeted gene testing was performed in cases of obvious immunodeficiency. For cases with less conclusive phenotypes, a gene panel was performed, followed by whole-exome sequencing if necessary.
Results
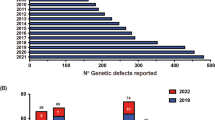
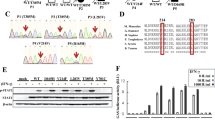
A total of 185 patients were diagnosed with IEIs during the study period; of these, 60.5% were male. Mean ages at symptom onset and diagnosis were 30.0 and 50.5 months, respectively. Consanguinity and a family history of IEIs were present in 86.9% and 50.8%, respectively. Most patients presented with lower respiratory infections (65.9%), followed by growth and development manifestations (43.2%). Phagocytic defects were the most common cause of IEIs (31.9%), followed by combined immunodeficiency (21.1%). Overall, 109 of 132 patients (82.6%) who underwent genetic testing received a genetic diagnosis, while testing was inconclusive for the remaining 23 patients (17.4%). Among patients with established diagnoses, 37 genes and 44 variants were identified. Autosomal recessive inheritance was present in 81.7% of patients with gene defects. Several variants were novel. Intravenous immunoglobulin therapy was administered to 39.4% of patients and 21.6% received hematopoietic stem cell transplantation. The overall survival rate was 75.1%.
Conclusion
This study highlights the genetic causes of IEIs in Omani patients. This information may help in the early identification and management of the disease, thereby improving survival and quality of life.


Similar content being viewed by others
Data Availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on request.
References
CEREDIH: The French PID study group. The French national registry of primary immunodeficiency diseases. Clin Immunol. 2010;135(2):264–72. https://doi.org/10.1016/j.clim.2010.02.021.
Bruton OC. Agammaglobulinemia Pediatrics. 1952;9(6):722–8. https://doi.org/10.1542/peds.9.6.722.
Kobrynski L, Powell RW, Bowen S. Prevalence and morbidity of primary immunodeficiency diseases, United States 2001–2007. J Clin Immunol. 2014;34(8):954–61. https://doi.org/10.1007/s10875-014-0102-8.
Wong M. What has happened in the last 50 years in immunology. J Paediatr Child Health. 2015;51(2):135–9. https://doi.org/10.1111/jpc.12834.
Fernandez Perez ER, Hunter M, Katial RK. United States trends in mortality rates for primary immunodeficiency diseases. J Allergy Clin Immunol Pract. 2019;7(3):1045–8. https://doi.org/10.1016/j.jaip.2018.09.030.
Odnoletkova I, Kindle G, Quinti I, Grimbacher B, Knerr V, Gathmann B, et al. The burden of common variable immunodeficiency disorders: a retrospective analysis of the European Society for Immunodeficiency (ESID) registry data. Orphanet J Rare Dis. 2018;13(1):201. https://doi.org/10.1186/s13023-018-0941-0.
Viti R, Marcellusi A, Capone A, Matucci A, Vultaggio A, Pignata C, et al. Direct and indirect costs of immunoglobulin replacement therapy in patients with common variable immunodeficiency (CVID) and X-linked agammaglobulinemia (XLA) in Italy. Clin Drug Investig. 2018;38(10):955–65. https://doi.org/10.1007/s40261-018-0688-3.
Modarresi SZ, Sabetkish N, Badalzadeh M, Tajik S, Esmaeili B, Fazlollahi MR, et al. The critical role of prenatal genetic study in prevention of primary immunodeficiency in high-risk families: the largest report of 107 cases. Iran J Allergy, Asthma, Immunol. 2020;19(5):478–83. https://doi.org/10.18502/ijaai.v19i5.4463.
Mateu L, Teniente-Serra A, Rocamora G, Marin-Muñiz A, Pàrraga N, Casas I, et al. Effect of an awareness campaign on the diagnosis and clinical impact of primary immunodeficiency. Med Clin (Barc). 2021;156(6):270–6. https://doi.org/10.1016/j.medcli.2020.04.066.
Al-Tamemi S, Naseem SU, Al-Siyabi N, El-Nour I, Al-Rawas A, Dennison D. Primary immunodeficiency diseases in Oman: 10-year experience in a tertiary care hospital. J Clin Immunol. 2016;36(8):785–92. https://doi.org/10.1007/s10875-016-0337-7.
Oliveira JB, Fleisher TA. Laboratory evaluation of primary immunodeficiencies. J Allergy Clin Immunol. 2010;125(2 Suppl 2):S297-305. https://doi.org/10.1016/j.jaci.2009.08.043.
Orange JS, Ballow M, Stiehm ER, Ballas ZK, Chinen J, De La Morena M, et al. Use and interpretation of diagnostic vaccination in primary immunodeficiency: a working group report of the Basic and Clinical Immunology Interest Section of the American Academy of Allergy, Asthma & Immunology. J Allergy Clin Immunol. 2012;130(3 Suppl):S1-24. https://doi.org/10.1016/j.jaci.2012.07.002.
Kanegane H, Hoshino A, Okano T, Yasumi T, Wada T, Takada H, et al. Flow cytometry-based diagnosis of primary immunodeficiency diseases. Allergol Int: Off J Japan Soc Allergol. 2018;67(1):43–54. https://doi.org/10.1016/j.alit.2017.06.003.
Salzer U, Sack U, Fuchs I. Flow cytometry in the diagnosis and follow up of human primary immunodeficiencies. EJIFCC. 2019;30(4):407–22.
Reda SM, El-Ghoneimy DH, Afifi HM. Clinical predictors of primary immunodeficiency diseases in children. Allergy Asthma Immunol Res. 2013;5(2):88–95. https://doi.org/10.4168/aair.2013.5.2.88.
Gathmann B, Mahlaoui N, Ceredih GL, Oksenhendler E, Warnatz K, et al. Clinical picture and treatment of 2212 patients with common variable immunodeficiency. J Allergy Clin Immunol. 2014;134(1):116–26. https://doi.org/10.1016/j.jaci.2013.12.1077.
Podjasek JC, Abraham RS. Autoimmune cytopenias in common variable immunodeficiency. Front Immunol. 2012;3:189. https://doi.org/10.3389/fimmu.2012.00189.
Todoric K, Koontz JB, Mattox D, Tarrant TK. Autoimmunity in immunodeficiency. Curr Allergy Asthma Rep. 2013;13(4):361–70. https://doi.org/10.1007/s11882-013-0350-3.
Hernandez-Trujillo VP, Scalchunes C, Cunningham-Rundles C, Ochs HD, Bonilla FA, Paris K, et al. Autoimmunity and inflammation in X-linked agammaglobulinemia. J Clin Immunol. 2014;34(6):627–32. https://doi.org/10.1007/s10875-014-0056-x.
Lehman HK. Autoimmunity and immune dysregulation in primary immune deficiency disorders. Curr Allergy Asthma Rep. 2015;15(9):53. https://doi.org/10.1007/s11882-015-0553-x.
Jonkman-Berk BM, van den Berg JM, Ten Berge IJ, Bredius RG, Driessen GJ, Dalm VA, et al. Primary immunodeficiencies in the Netherlands: national patient data demonstrate the increased risk of malignancy. Clin Immunol. 2015;156(2):154–62. https://doi.org/10.1016/j.clim.2014.10.003.
Kersey JH, Spector BD, Good RA. Primary immunodeficiency diseases and cancer: the immunodeficiency-cancer registry. Int J Cancer. 1973;12(2):333–47. https://doi.org/10.1002/ijc.2910120204.
Tiri A, Masetti R, Conti F, Tignanelli A, Turrini E, Bertolini P, et al. Inborn errors of immunity and cancer. Biology (Basel). 2021;10(4):313. https://doi.org/10.3390/biology10040313.
Kwan A, Abraham RS, Currier R, Brower A, Andruszewski K, Abbott JK, et al. Newborn screening for severe combined immunodeficiency in 11 screening programs in the United States. JAMA. 2014;312(7):729–38. https://doi.org/10.1001/jama.2014.9132.
El-Sayed ZA, Radwan N. Newborn screening for primary immunodeficiencies: the gaps, challenges, and outlook for developing countries. Front Immunol. 2019;10:2987. https://doi.org/10.3389/fimmu.2019.02987.
Hale JE, Platt CD, Bonilla FA, Hay BN, Sullivan JL, Johnston AM, et al. Ten years of newborn screening for severe combined immunodeficiency (SCID) in Massachusetts. J Allergy Clin Immunolo Pract. 2021;9(5):2060-7.e2. https://doi.org/10.1016/j.jaip.2021.02.006.
Maffucci P, Filion CA, Boisson B, Itan Y, Shang L, Casanova J-L, et al. Genetic diagnosis using whole exome sequencing in common variable immunodeficiency. Front Immunol. 2016;7:220. https://doi.org/10.3389/fimmu.2016.00220.
Simon AJ, Golan AC, Lev A, Stauber T, Barel O, Somekh I, et al. Whole exome sequencing (WES) approach for diagnosing primary immunodeficiencies (PIDs) in a highly consanguineous community. Clin Immunol. 2020;214: 108376. https://doi.org/10.1016/j.clim.2020.108376.
Rae W, Ward D, Mattocks C, Pengelly RJ, Eren E, Patel SV, et al. Clinical efficacy of a next-generation sequencing gene panel for primary immunodeficiency diagnostics. Clin Genet. 2018;93(3):647–55. https://doi.org/10.1111/cge.13163.
Arts P, Simons A, AlZahrani MS, Yilmaz E, AlIdrissi E, van Aerde KJ, et al. Exome sequencing in routine diagnostics: a generic test for 254 patients with primary immunodeficiencies. Genome Med. 2019;11(1):38. https://doi.org/10.1186/s13073-019-0649-3.
Cifaldi C, Brigida I, Barzaghi F, Zoccolillo M, Ferradini V, Petricone D, et al. Targeted NGS platforms for genetic screening and gene discovery in primary immunodeficiencies. Front Immunol. 2019;10:316. https://doi.org/10.3389/fimmu.2019.00316.
Arunachalam AK, Maddali M, Aboobacker FN, Korula A, George B, Mathews V, et al. Primary immunodeficiencies in India: molecular diagnosis and the role of next-generation sequencing. J Clin Immunol. 2021;41(2):393–413. https://doi.org/10.1007/s10875-020-00923-2.
Bousfiha A, Jeddane L, Picard C, Al-Herz W, Ailal F, Chatila T, et al. Human inborn errors of immunity: 2019 Update of the IUIS phenotypical classification. J Clin Immunol. 2020;40(1):66–81. https://doi.org/10.1007/s10875-020-00758-x.
Picard C, Bobby Gaspar H, Al-Herz W, Bousfiha A, Casanova JL, Chatila T, et al. International Union of Immunological Societies: 2017 primary immunodeficiency diseases committee report on inborn errors of immunity. J Clin Immunol. 2018;38(1):96–128. https://doi.org/10.1007/s10875-017-0464-9.
Badran YR, Massaad MJ, Bainter W, Cangemi B, Naseem SU, Javad H, et al. Combined immunodeficiency due to a homozygous mutation in ORAI1 that deletes the C-terminus that interacts with STIM 1. Clin Immunol. 2016;166–167:100–2. https://doi.org/10.1016/j.clim.2016.03.012.
Al-Tamemi S, Alhinai Z, Al-Rahbi N, Al-Abdawani R, Al-Yazidi L, Al-Shekaili J, et al. BCL10 loss-of-function novel mutation leading to atypical severe combined immunodeficiency. Clin Immunol. 2022;241:109067. https://doi.org/10.1016/j.clim.2022.109067.
Bainter W, Platt CD, Park SY, Stafstrom K, Wallace JG, Peters ZT, et al. Combined immunodeficiency due to a mutation in the γ1 subunit of the coat protein I complex. J Clin Invest. 2021;131(3): e140494. https://doi.org/10.1172/jci140494.
Engelhardt KR, McGhee S, Winkler S, Sassi A, Woellner C, Lopez-Herrera G, et al. Large deletions and point mutations involving the dedicator of cytokinesis 8 (DOCK8) in the autosomal-recessive form of hyper-IgE syndrome. J Allergy Clin Immunol. 2009;124(6):1289-302.e4.
Al-Marhoobi R, Al-Musalhi M, Wali Y, Alsayegh A, Al-Tamemi S. Combined immunodeficiency, hemolytic anemia, and growth retardation secondary to a homozygous mutation in the NHEJ1 gene. J Pediatr Hematol Oncol. 2020;42(4):333–5. https://doi.org/10.1097/MPH.0000000000001545.
Al-Zadjali S, Al-Tamemi S, Elnour I, AlKindi S, Lapoumeroulie C, Al-Maamari S, et al. Clinical and molecular findings of chronic granulomatous disease in Oman: family studies. Clin Genet. 2015;87(2):185–9. https://doi.org/10.1111/cge.12351.
Al-Riyami AZ, Al-Zadjali S, Al-Mamari S, Al-Said B, Al-Qassabi J, Al-Tamemi S. Correlation between flow cytometry and molecular findings in autosomal recessive chronic granulomatous disease: a cohort study from Oman. Int J Lab Hematol. 2018;40(5):592–6. https://doi.org/10.1111/ijlh.12873.
Nazir HF, Rawas AA, Tamemi SA, Zadjali SA, Hosni SA, Tauro M, et al. Hematopoietic stem cell transplantation for patients with autosomal recessive complete INF-λ receptor 2 deficiency: experience in Oman. Transplant Cell Ther. 2021;27(10):881.e1-5. https://doi.org/10.1016/j.jtct.2021.07.013.
Al-Tamemi S, Al-Zadjali S, Al-Ghafri F, Dennison D. Chediak-Higashi syndrome: novel mutation of the CHS1/LYST gene in 3 Omani patients. J Pediatr Hematol Oncol. 2014;36(4):e248–50. https://doi.org/10.1097/MPH.0000000000000025.
Platt CD, Zaman F, Bainter W, Stafstrom K, Almutairi A, Reigle M, et al. Efficacy and economics of targeted panel versus whole-exome sequencing in 878 patients with suspected primary immunodeficiency. J Allergy Clin Immunol. 2021;147(2):723–6. https://doi.org/10.1016/j.jaci.2020.08.022.
Al-Herz W, Chou J, Delmonte OM, Massaad MJ, Bainter W, Castagnoli R, et al. Comprehensive genetic results for primary immunodeficiency disorders in a highly consanguineous population. Front Immunol. 2018;9:3146. https://doi.org/10.3389/fimmu.2018.03146.
Engelbrecht C, Urban M, Schoeman M, Paarwater B, van Coller A, Abraham DR, et al. Clinical utility of whole exome sequencing and targeted panels for the identification of inborn errors of immunity in a resource-constrained setting. Front Immunol. 2021;12:665621. https://doi.org/10.3389/fimmu.2021.12:665621.
Aghamohammadi A, Rezaei N, Yazdani R, Delavari S, Kutukculer N, Topyildiz E, et al. Consensus Middle East and North Africa registry on inborn errors of immunity. J Clin Immunol. 2021;41(6):1339–51. https://doi.org/10.1007/s10875-021-01053-z.
Köker MY, Camcıoğlu Y, van Leeuwen K, Kılıç S, Barlan I, Yılmaz M, et al. Clinical, functional, and genetic characterization of chronic granulomatous disease in 89 Turkish patients. J Allergy Clin Immunol. 2013;132(5):1156-63.e5. https://doi.org/10.1016/j.jaci.2013.05.039.
de Boer M, Tzur S, van Leeuwen K, Dencher PC, Skorecki K, Wolach B, et al. A founder effect for p47(phox)Trp193Ter chronic granulomatous disease in Kavkazi Jews. Blood Cells Mol Dis. 2015;55(4):320–7. https://doi.org/10.1016/j.bcmd.2015.07.014.
Islam MM. The practice of consanguineous marriage in Oman: prevalence, trends and determinants. J Biosoc Sci. 2012;44(5):571–94. https://doi.org/10.1017/S0021932012000016.
Rezaei N, Pourpak Z, Aghamohammadi A, Farhoudi A, Movahedi M, Gharagozlou M, et al. Consanguinity in primary immunodeficiency disorders; the report from Iranian Primary Immunodeficiency Registry. Am J Reprod Immunol. 2006;56(2):145–51. https://doi.org/10.1111/j.1600-0897.2006.00409.x.
Abd Elaziz D, Abd El-Ghany M, Meshaal S, El Hawary R, Lotfy S, Galal N, et al. Fungal infections in primary immunodeficiency diseases. Clin Immunol. 2020;219: 108553. https://doi.org/10.1016/j.clim.2020.108553.
Demirdag YY, Gupta S. Update on infections in primary antibody deficiencies. Front Immunol. 2021;12: 634181. https://doi.org/10.3389/fimmu.2021.634181.
Khalili N, Mohammadzadeh I, Khalili N, Heredia RJ, Zoghi S, Boztug K, et al. BCGitis as the primary manifestation of chronic granulomatous disease. IDCases. 2020;23: e01038. https://doi.org/10.1016/j.idcr.2020.e01038.
Kuruvilla M, de la Morena MT. Antibiotic prophylaxis in primary immune deficiency disorders. J Allergy Clin Immunol Pract. 2013;1(6):573–82. https://doi.org/10.1016/j.jaip.2013.09.013.
Pai SY, Logan BR, Griffith LM, Buckley RH, Parrott RE, Dvorak CC, et al. Transplantation outcomes for severe combined immunodeficiency, 2000–2009. N Engl J Med. 2014;371(5):434–46. https://doi.org/10.1056/NEJMoa1401177.
Chan AY, Leiding JW, Liu X, Logan BR, Burroughs LM, Allenspach EJ, et al. Hematopoietic cell transplantation in patients with primary immune regulatory disorders (PIRD): a Primary Immune Deficiency Treatment Consortium (PIDTC) survey. Front Immunol. 2020;11:239. https://doi.org/10.3389/fimmu.2020.00239.
Al-Herz W. Mortality among primary immunodeficient patients in Kuwait. J Allergy Clin Immunol. 2010;125(2S1):AB75. https://doi.org/10.1016/j.jaci.2009.12.296.
Sasaki K, Jabbour E, Short NJ, Jain N, Ravandi F, Pui CH, et al. Acute lymphoblastic leukemia: a population-based study of outcome in the United States based on the Surveillance, Epidemiology, and End Results (SEER) database, 1980–2017. Am J Hematol. 2021;96(6):650–8. https://doi.org/10.1097/MPH.0000000000001545.
Acknowledgements
We would like to thank Prof. Raif Geha, Harvard University for his valuable review of the manuscript. We would like to thank all staff who participated in the care of these patients and all patients and their families for their trust.
Funding
Part of this study was funded by a Sultan Qaboos University research grant. No support was received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
The corresponding author was responsible for all aspects of the study conception and design as well as the preparation of the manuscript for publication. SZ & MR performed targeted gene sequencing and analysis; DD supervised the targeted gene sequence and analysis; SN and NS participated in clinical care and family support; KA performed genetic counseling; ZB, KD, FM, AS, and AM supervised panel gene testing and analysis of panel and whole-exome sequencing.
Corresponding author
Ethics declarations
Ethics Approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Medical Research and Ethics Committee of the College of Medicine and Health Sciences, Sultan Qaboos University, Muscat, Oman.
Consent to Participate
Informed consent was obtained from all patients prior to genetic testing.
Consent for Publication
The authors affirm that no identifying participant data was published in any form.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Al-Tamemi, S., Al-Zadjali, S., Bruwer, Z. et al. Genetic Causes, Clinical Features, and Survival of Underlying Inborn Errors of Immunity in Omani Patients: a Single-Center Study. J Clin Immunol 43, 452–465 (2023). https://doi.org/10.1007/s10875-022-01394-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10875-022-01394-3